Metabolomics
1. Can you list at least FOUR classes of molecules as subjects for metabolomics studies?
- Peptides
- Oligonucleotides
- Sugars
- Nucleosides
- Organic acids
- Ketones
- Aldehydes
- Amines
- Amino acids
- Lipids
- Steroids
- Alkaloids
- Drugs (xenobiotics)
2. What solvents do you commonly use to extract metabolites in metabolomic studies with complex samples?
Aqueous or methanolic solvents.
3. GC-MS and NMR are two of the most important tools for metabolomics studies. Please list at least two advantages and two disadvantages of GC-MS compared to NMR studies.
Advantages:
- Powerful technique: Wide range of applications from identification to quantification of a variety of compounds in a sample.
- Highly sensitive method: MS can readily quantify very low molecular concentrations, down to nanomolar and picomolar.
- A large number of metabolites: When coupled with chromatography, both liquid chromatography (LC) and gas chromatography (GC), MS can detect hundreds and thousands of metabolites in a given sample.
- Analysis of secondary metabolites: Chromatography, when coupled with MS, also allows one to analyse secondary metabolites even where the detection level is lower.
- Small sample volume: When using the MS technology, it is possible to measure sample volumes as low as 10 ul serum/plasma.
Disadvantages:
- Batch effect: When chromatography is coupled with MS, the sample unavoidably interacts directly with the instrument. This, over time, leads to changes in measured analyte response.
- Moderate reproducibility: Since the sample interacts with the instrument, it decreases its precision levels.
- Sample recovery: While NMR allows using the same sample for multiple analysis, in MS, once a sample is analysed it can’t be recovered.
- Slow analysis: Depending on the protocol used, analysis time can range from a few minutes and go up to one hour per sample, which makes it significantly slower than NMR.
- Expensive analysis: MS requires sample preparation and separation (reagents as well as standards for metabolite identification), which makes it a costlier technique than NMR.
4. In the case study with targeted metabolomics, what is the purpose of adding etodolac into samples? Can you write down the transition of 286.1 to 212.1?
As internal standard.
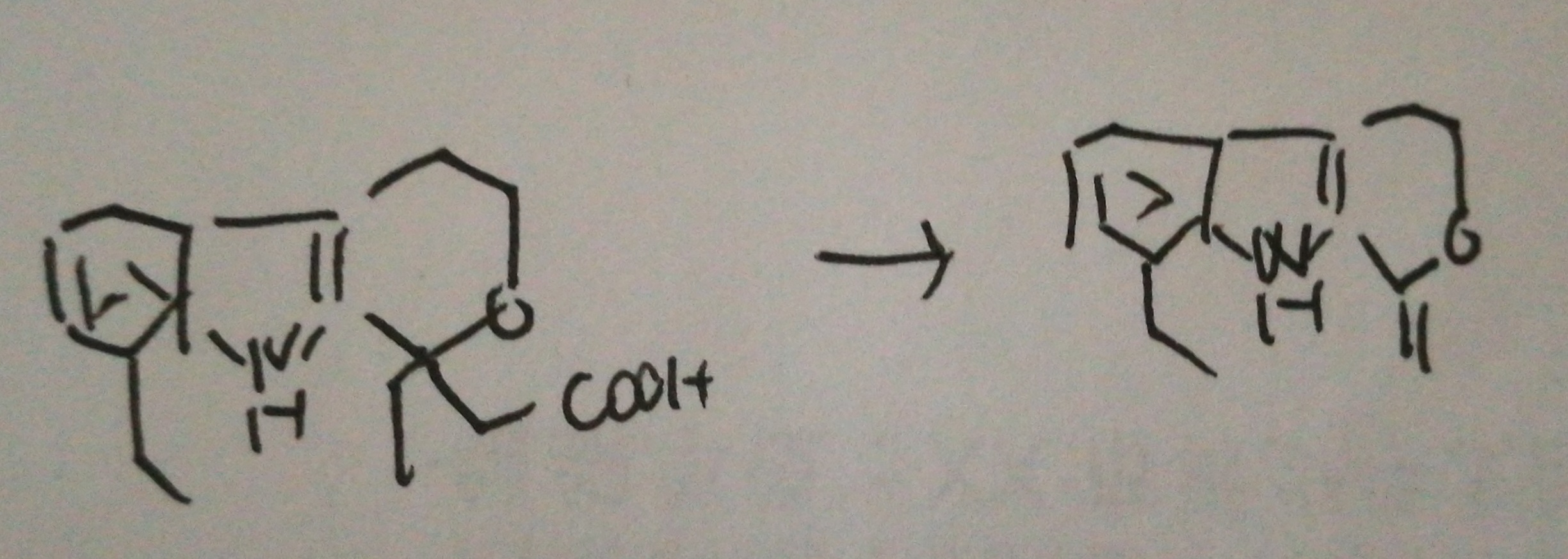
5. In untargeted metabolomics, one challenge is to identify compounds in the complex metabolomic samples. From the studies shown by Bruce Cooper, 6 hits were identified through database search. How do you verify their IDs?
Using authentic standards for retention time match.